Journal: bioRxiv
Article Title: Phosphoribosyl ubiquitination of SNARE proteins regulate autophagy in Legionella infection
doi: 10.1101/2025.05.19.654886
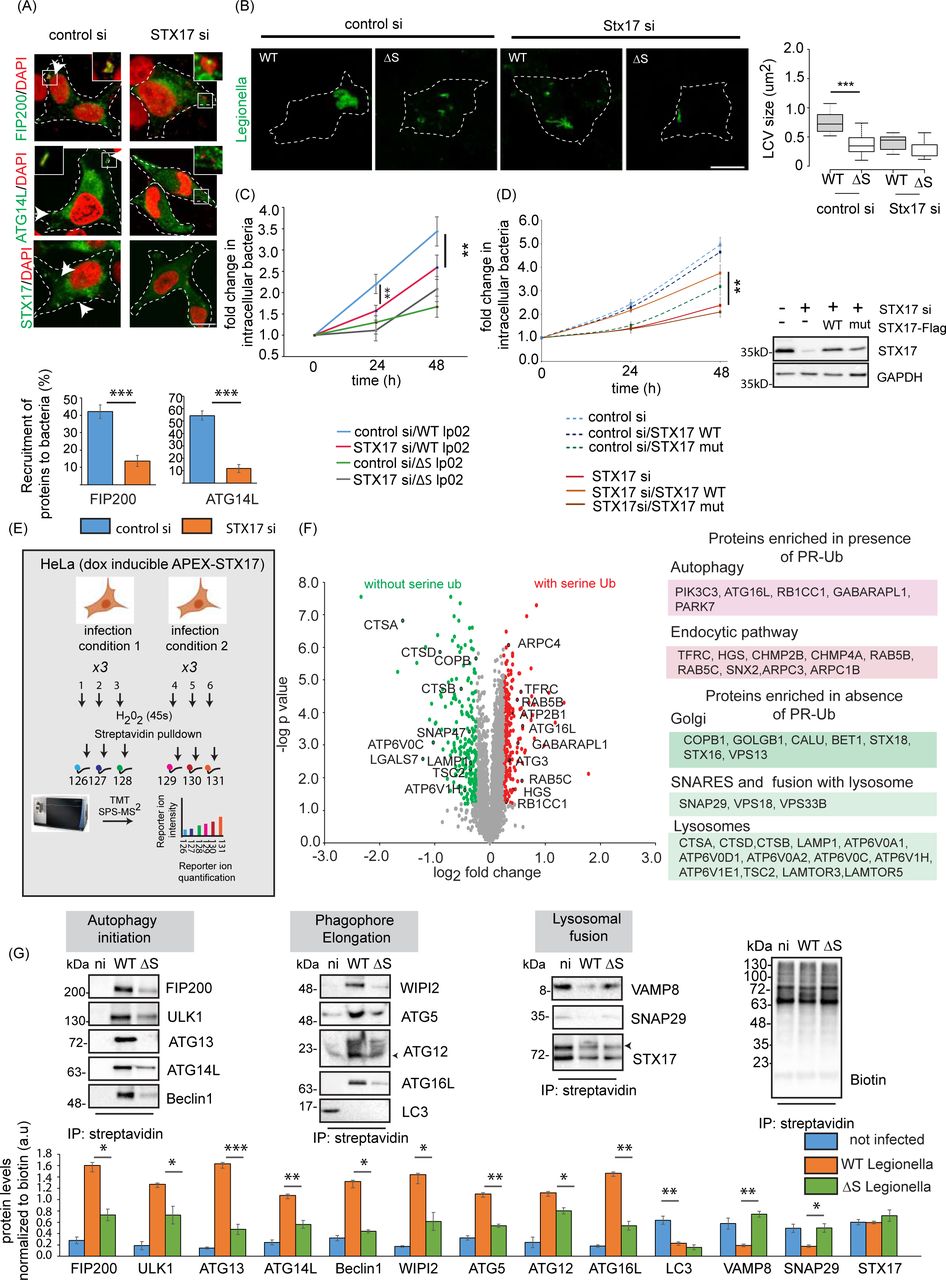
Figure Lengend Snippet: a) A549 cells were treated with control or STX17 siRNA for 48 h followed by infection with WT Legionella . Cells were fixed and stained for indicated autophagy markers 1 h after infection. The data are means ± SEM of 50 cells representing three experiments. which were analyzed per sample to quantitate recruitment of FIP200 and ATG14L to bacteria. p value was calculated using 2 tailed type 3 Student’s t-test, ***p=5.83E-7(FIP200), ***p=1.92E-12(ATG14L), Scale bar: 5µm. Dotted lines indicate cell outlines drawn from thresholding images in FIJI. b) A549 cells were treated with STX17 or control siRNA for 48 h followed by infection with WT or ΔS Legionella for 12 h (MOI = 1). Cells were fixed for immunostaining with a Legionella -specific antibody followed by confocal microscopy. The LCV size was estimated in FIJI. In the box-plot, center lines show the medians; box limits indicate the 25th and 75th percentiles as determined by R software; whiskers extend 1.5 times the interquartile range from the 25th and 75th percentiles, outliers are represented by dots. n = 32, 31 cells taken from 3 independent experiments. p value was calculated using 2 tailed, type 3 Student’s t-test, ***p =8.13E-16. Scale bar: 5µm. Dotted lines indicate cell outlines drawn from thresholding images in FIJI. c) A549 cells were treated with control or STX17 siRNA for 48 h followed by infection with WT or ΔS Legionella . Intracellular bacterial replication was assessed after 0, 24 and 48 h. Data are means ± SEM of three independent experiments. p value was calculated using 2 tailed type 3 Student’s t-test, **p =0.00526 (WT, control vs STX17siRNA, 24h), **p =0.00815 (WT, control vs STX17siRNA, 48h), d) A549 cells were treated with STX17 or control siRNA for 48 h followed by transfection with WT or PR-Ub-deficient STX17 for 24 h. Intracellular bacterial replication was assessed after 0, 24 and 48 h. Data are means ± SEM of three independent experiments **p=0.0077 (WT, STX17siRNA versus STX17 mutant, STX17siRNA, 48 h).Western blotting with STX17 antibody shows knockdown efficiency of STX17 siRNA and reconstitution with WT or mutant STX17. e) Proximity labeling assay workflow. HeLa cells expressing doxycycline-inducible APEX-STX17 and CD32(for increasing efficiency of Legionella uptake) were infected with Legionella for 2 h before treatment with biotin-tyramide and H 2 O 2 followed by streptavidin pulldown. The samples were reduced, alkylated and digested with trypsin before MS analysis. Samples representing three biological replicates each of non-infected and Legionella -infected cells were analyzed in a single reaction by 6plex TMT labeling. f) Volcano plot showing changes in the biotin-labeled proteome following the infection of HeLa cells expressing APEX2-FLAG-STX17 with ΔR and ΔRΔS Legionella for 2 h, GO analysis of the biotin-labeled proteome showing pathways upregulated by infection with ΔR vs ΔRΔS Legionella . Data represents mean fold change of three experimental replicates per infection set (n=3). p value was calculated using 2 tailed type 3 Student’s t-test and significant candidates were chosen having p-value ≤0.01 and log2(fold change) value minimum of ±0.5. Red and green indicate compartments containing proteins enriched following infection with ΔR and ΔRΔS Legionella , respectively. g) Cells expressing doxycycline-inducible APEX-STX17 were infected with Legionella for 2 h before treatment with biotin-tyramide and H 2 O 2 followed by streptavidin pulldown. The samples were analyzed by western blot with antibodies against proteins of the autophagic and endosomal pathways. The data represents means ± SD of 3 independent experiments. p value was calculated using 2 tailed type 3 Students t-test. WT vs ΔS p values: *p= 0.01006 (FIP200), **p=0.0206 (ULK1), p=***0.0007 (ATG13) **p=0.005 (ATG14), *p=0.0111(Beclin1), *p=0.0219 (WIPI2), **p=0.0038 (ATG5), *p=0.018 (ATG12), **p=0.001 (ATG16), **p=0.0036 (VAMP8), *p=0.0424 (SNAP29). (n.i-not infected, WT-wild-type Legionella , ΔS-ΔSidE Legionella )
Article Snippet: We used the following antibodies and dilutions: STX17 (cat. no. 17815-1-AP, Proteintech; 1:1000), GAPDH (cat. no. 2118, Cell Signaling Technology; 1:2000), GFP trap beads (cat. no. gta-100, ChromoTek), GFP (cat. no. sc-9996, Santa Cruz Biotechnology; 1:2000), GFP for immune-electron microscopy (cat no. 600–106-215, Rockland), biotin for immuno-electron microscopy (cat. no. 100–4198, Rockland), LC3 (cat. no. 2775, Cell Signaling Technology; 1:2000), VAMP8 (cat. no. 13060, Cell Signaling Technology; 1:1000), SNAP29 (cat. no. 3013, Cell Signaling Technology; 1:2000), RAB5 (cat. no. 3547; 1:1,000), RAB7 (cat. no. 9367, Cell Signaling Technology; 1:2000), ATG16L (cat. no. ab188642, Cell Signaling Technology; 1:1000), ATG12 (cat. no. 4180, Cell Signaling Technology; 1:1000), Beclin1 (cat. no. 3738, Cell Signaling Technology; 1:1000), FIP200 (cat. no. 17250-1AP, Proteintech; 1:1000), LAMP1 (cat. no. 9091, Cell Signaling Technology; 1:2000), and Legionella (cat. no. 20943, Abcam; 1:4000), Ubiquitin (Cat. no: 3933, Cell Signaling Technology, 1:1000).
Techniques: Control, Infection, Staining, Bacteria, Immunostaining, Confocal Microscopy, Software, Transfection, Mutagenesis, Western Blot, Knockdown, Labeling, Expressing